Isozymes (also known as isoenzymes) are enzymes that differ in amino acid sequence but catalyze the same chemical reaction. These enzymes usually display different kinetic parameters (i.e. different KM values), or different regulatory properties. The existence of isozymes permits the fine-tuning of metabolism to meet the particular needs of a given tissue or developmental stage (for example lactate dehydrogenase (LDH)). In biochemistry, isozymes (or isoenzymes) are isoforms (closely related variants) of enzymes. In many cases, they are coded for by homologous genes that have diverged over time. Although, strictly speaking, allozymes represent different alleles of the same gene, and isozymes represent different genes whose products catalyse the same reaction, the two words are usually used interchangeably.
Isozymes were first described by hunter and Markert (1957) who defined them as different variants of the same enzyme having identical functions and present in the same individual. This definition encompasses (1) enzyme variants that are the product of different genes and thus represent different loci (described as isozymes) and (2) enzymes that are the product of different alleles of the same gene (described as allozymes).
Isozymes are usually the result of gene duplication, but can also arise from polyploidisation or hybridization. Over evolutionary time, if the function of the new variant remains identical to the original, then it is likely that one or the other will be lost as mutations accumulate, resulting in a pseudogene. However, if the mutations do not immediately prevent the enzyme from functioning, but instead modify either its function, or its pattern of gene Expression, then the two variants may both be favoured by natural selection and become specialised to different functions. For example, they may be expressed at different stages of development or in different tissues.
Allozymes may result from point mutations or from insertion-deletion (indel) events that affect the dna coding sequence of the gene. As with any other new mutation, there are three things that may happen to a new allozyme:
It is most likely that the new allele will be non-functional — in which case it will probably result in low fitness and be removed from the population by natural selection.
Alternatively, if the amino acid residue that is changed is in a relatively unimportant part of the enzyme, for example a long way from the active site then the mutation may be selectively neutral and subject to genetic drift.
In rare cases the mutation may result in an enzyme that is more efficient, or one that can catalyse a slightly different chemical reaction, in which case the mutation may cause an increase in fitness, and be favoured by natural selection.
An example of an isozyme:
An example of an isozyme is glucokinase, a variant of hexokinase which is not inhibited by glucose 6-phosphate. Its different regulatory features and lower affinity for glucose (compared to other hexokinases), allows it to serve different functions in cells of specific organs, such as control of insulin release by the beta cells of the pancreas, or initiation of glycogen synthesis by liver cells. Both of these processes must only occur when glucose is abundant, or problems occur.
Dictionary > Isoenzyme
You will also like...
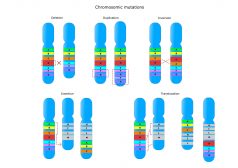

Chromosome Mutations
Mutations can also influence the phenotype of an organism. This tutorial looks at the effects of chromosomal mutations, ..
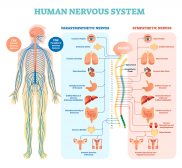
The Human Nervous System
The nervous system is essentially a biological information highway. This tutorial gives an overview of the nervous syste..

Plant Water Regulation
Plants need to regulate water in order to stay upright and structurally stable. Find out the different evolutionary adap..
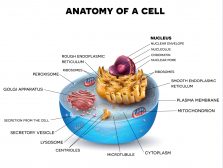
Biological Cell Introduction
It only takes one biological cell to create an organism. A single cell is able to keep itself functional through its 'mi..

Biosecurity and Biocontrol
This lesson explores the impact of biosecurity threats, and why they need to be identified and managed. Examples to incl..

Still Water Animals
Animals living in aquatic habitats have diversified and evolved through time. They eventually occupy ecological niches a..

